A state-of-the-art analysis tool for calcium imaging
For the annotation, downstream analysis and curation of calcium imaging data recorded by Femtonics microscopes, we highly recommend the Mesmerize package written by Kushal Kolar from the Chatzigeorgiou Lab.

Mesmerize encompasses the entire process of calcium imaging analysis, from raw data to interactive, semi-final publication figures, and aids users in creating FAIR-functionally linked datasets. It is applicable to a broad range of experiments and is intended for users with or without a programming background.
Mesmerize can import MESc and MES files and contains front-end GUI modules for
the latest cutting-edge techniques for data pre-processing, signal extraction,
and analysis. Mesmerize integrates with the CaImAn library,
which implements fast and scalable algorithms for motion correction, automatic
source extraction, and spike deconvolution. Mesmerize also provides the option
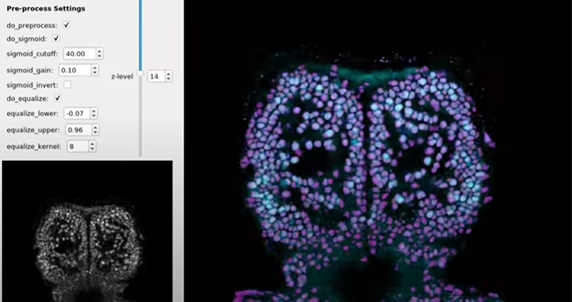
of using deep-learning for cell segmentation, which is more suitable for
nuclear localized indicators.
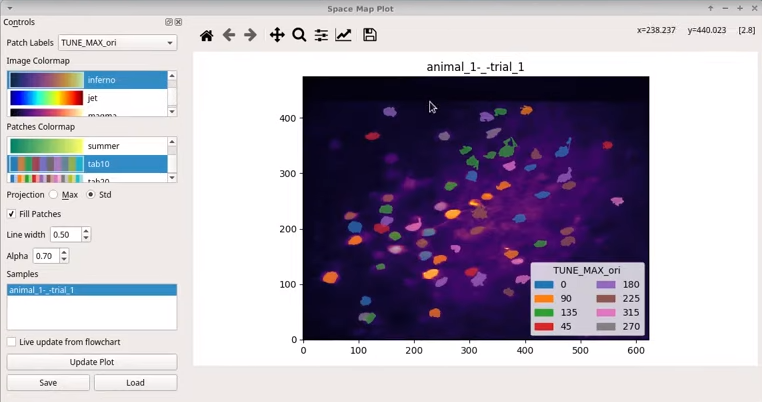
The Mesmerize flowchart allows users to quickly perform downstream using a GUI.
You can perform signal processing, create stimulus tuning curves, perform
various types of clustering analysis and create interactive plots, which store
the analysis procedures to aid reproducibility.
Watch a video about a new deep learning method. It’s very fast, requires litter parameter tuning and it only takes a few minutes to segment 30 planes of a zebrafish.
Watch the Mesmerize tutorials
Check out MES and MESc importers
We are delighted that you are interested in Mesmerize. In case you have any questions regarding the installation and the utilization of the software you can find the relevant information on GitHub. Femtonics shall not be liable for the software as a product, however endorses the use of Kushal Kolar’s and Femtonics’ software developers’ intellectual property.